Quantitative Imaging of the Cell
Team Leader : Jean-Baptiste Sibarita
General objective

General objectives
We develop cutting-edge quantitative imaging methods to decipher protein organization and dynamics with high spatial and temporal resolution and compatible with high-content screening standards. Over the last 10 years, we have successfully developed and combined single-molecule based nanoscopy, dedicated image analysis methods and bioengineering techniques to tackle important biological questions. With no relevant biological question to address within the team itself, our developments are applied in very close collaboration with biology research groups.
Our team has an important academic and industrial technology transfer activity. We develop software and microscopy solutions which we valorize through scientific publications, patents, industrial technology transfers, academic Material Transfer Agreements, and free/collaborative/open source distribution.
Team organization
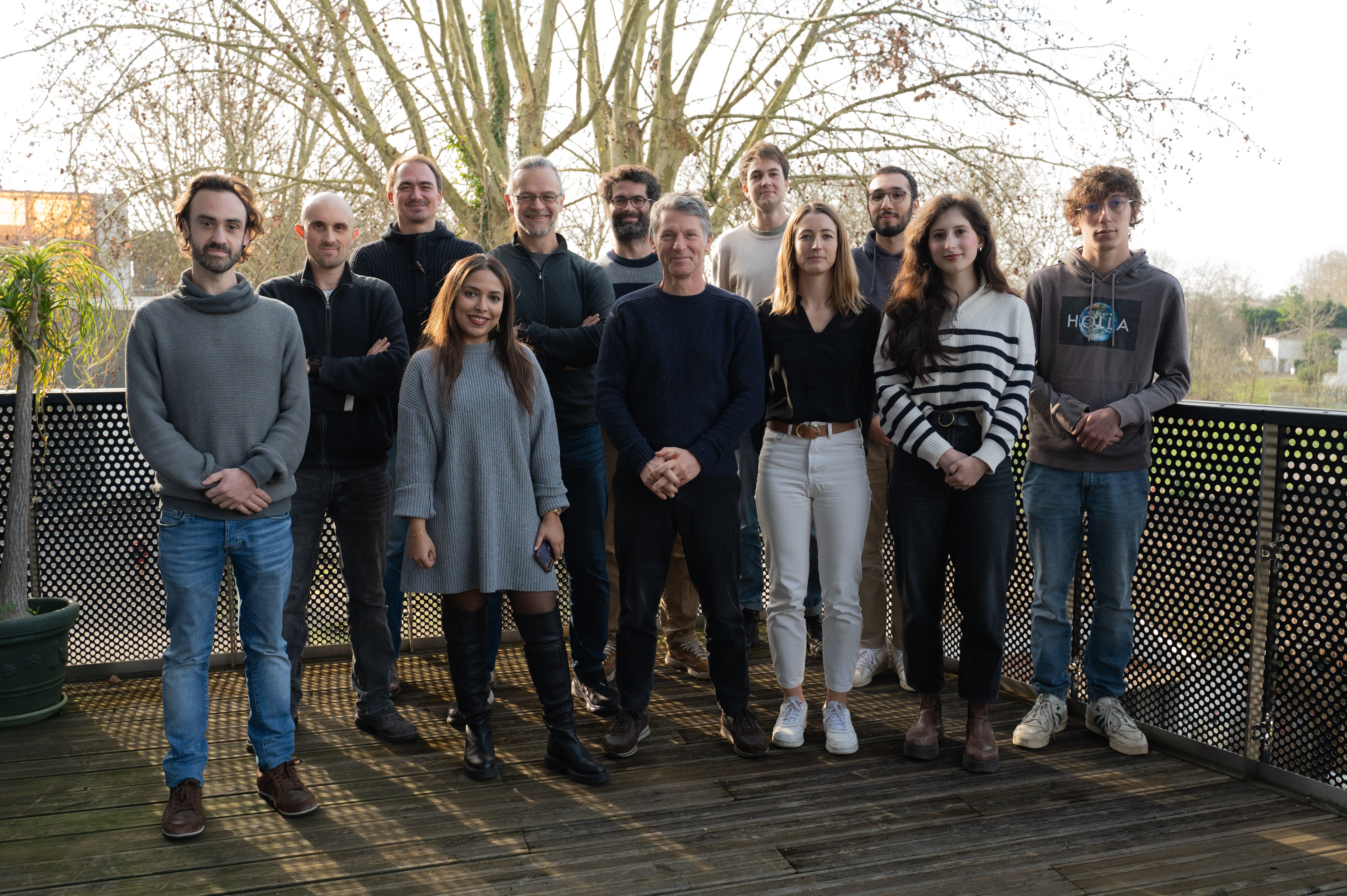
The Quantitative imaging of the cell team is a R&D team composed of people coming from various disciplines: microscopy, computer science and bioengineering.
Project leaders
 |
 |
 |
|
Jean-Baptiste Sibarita |
Florian Levet |
Rémi Galland |
Research Projects
3D high- and super-resolution imaging at different scales
MORE

Quantitative analysis of super-resolution data
MORE

Single-molecule-based High Content Screening
MORE

Correlative platform for SMLM & STED nanoscopy
MORE

High-content screening imaging of living organoids
MORE
Combining deep-learning with geometric and image processing for analysis of microscopy data
MORE
PalmTracer
MOREExpertise
News
A framework for evaluating the performance of SMLM cluster analysis algorithms - Nature, Feb 2023
Daniel J. Nieves, Jeremy A. Pike, Florian Levet, David J. Williamson, Mohammed Baragilly, Sandra Oloketuyi, Ario de Marco, Juliette Griffie, Daniel Sage, Edward A. K. Cohen, Jean-Baptiste Sibarita, Mike Heilemann, Dylan M. Owen
Nat Methods. 2023-02-01; 20(2): 259-267
DOI: 10.1038/s41592-022-01750-6
Single-molecule localization microscopy (SMLM) generates data in the form of coordinates of localized fluorophores. Cluster analysis is an attractive route for extracting biologically meaningful information from such data and has been widely applied. Despite a range of cluster analysis algorithms, there exists no consensus framework for the evaluation of their performance. Here, we use a systematic approach based on two metrics to score the success of clustering algorithms in simulated conditions mimicking experimental data. We demonstrate the framework using seven diverse analysis algorithms: DBSCAN, ToMATo, KDE, FOCAL, CAML, ClusterViSu and SR-Tesseler. Given that the best performer depended on the underlying distribution of localizations, we demonstrate an analysis pipeline based on statistical similarity measures that enables the selection of the most appropriate algorithm, and the optimized analysis parameters for real SMLM data. We propose that these standard simulated conditions, metrics and analysis pipeline become the basis for future analysis algorithm development and evaluation.
The point of view of Jean-Baptiste Sibarita and Florian Levet (co-authors)
This work by Nieves et al. proposes metrics to evaluate the performances of clustering methods dedicated to single molecule localization microscopy (SMLM) data. It includes the methods and several reference simulation datasets to assess the performances of existing and future approaches. This is a consortium paper that regroups word wide recognized experts in the field of SMLM. Amongst them, Florian Levet and Jean-Baptiste Sibarita from the Quantitative imaging of the Cell team (IINS) & Bordeaux Imaging Center, contributed to this work as developers of SR-Tesseler and Coloc-Tesseler software and the corresponding methods published in (Levet et al, Nature Methods 2015) and (Levet et al, Nature Comm, 2019).
PoCA: a software platform for point cloud data visualization and quantification, Nature - March 2023
Levet, F., Sibarita, JB
Nat Methods. 2023-03-06
https://doi.org/10.1038/s41592-023-01811-4
Point Cloud Analyst (PoCA) is a powerful open-source software platform dedicated to the visualization and analysis of 2D and 3D point-cloud data. PoCA allows manipulating large datasets, and integrates a plugin architecture, a native batch analysis engine and a Python code interpreter, facilitating both the analysis of data and the integration of new methods.
PoCA is packaged as a one-click installer for the Windows operating system. The source code is available on GitHub (https://github.com/flevet/PoCA) under a LGPL v3 license. We provide a cmake script to facilitate its building inside other operating systems. The documentation is available at https://poca-smlm.github.io/.
To know more:
Job offer: Engineer to develop light-sheet microscopy for neurosciences applications
The Sibarita team "Quantitative Imaging of the Cell" is seeking an engineer to develop light-sheet microscopy for neurosciences applications.
Project description
Light Sheet Fluorescence Microscopy technics (LSFM) have proven to be extremely efficient for 3D imaging of biological samples at various spatial and temporal scales with minimal photo-damaging effects. Several solutions have been developed in the field of neuroscience to image samples ranging from fixed whole brains, to single dissociated neurons growing on a coverslip. In this regard, the Interdisciplinary Institute for Neuroscience (IINS) and the Bordeaux Imaging Center (BIC) are equipped with 3 complementary LSFM techniques: (1) an ultramicroscope for whole brain imging; (2) a Lattice
Light Sheet Microscope (LLSM) to image the first layers of brain slices at high spatial resolution; (3) a home-made single objective selective plane illumination microscope (soSPIM) dedicated to 3D cell cultures and in-depth single-molecule localization microscopy (SMLM).
We aim to complete our catalog by implementing a solution based on the Oblique Plane Microscopy (OPM) architecture, which will be dedicated to fast neuronal samples imaging, ie. brain slices, equipped with local photo-manipulation and possibly other recording modalities.
Missions
The candidate missions will be to:
- develop a custom OPM to address specific neurobiological questions
- participate to the improvement of existing light-sheet microscopes in close collaboration with developers and neuroscientists.
Candidate profile
We seek a motivated, enthusiastic and independent candidate, with an interest in neuroscience and a strong expertise in optics and fluorescence microscopy. Complementary skills in programming and sample preparation are appreciated. The candidate will work in an English-speaking environment, in close interactions with the neuroscientists' team of the Bordeaux's Neurocampus.
Contract
A 3 years research engineer position is available in the framework of the French "Grands Programmes de Recherche" BRAIN awarded to the Bordeaux Neurocampus.
Applicants should send a CV, a motivation letter and contact details for at least two referees to:
jean-baptiste.sibarita@u-bordeaux.fr; remi.galland@u-bordeaux.fr; mathieu.ducros@u-bordeaux.fr
Job offer: Engineer to automated fluidics and microfabrication for multi-conditions 3D biological
The Sibarita team "Quantitative Imaging of the Cell" is seeking an engineer to develop automated fluidics and microfabrication approaches for multi-conditions observation of 3D biological samples using the soSPIM technology.
Project description
Spheroids and organoids have emerged in the last decade as very promising biological models for applications ranging from fundamental research to toxicology assays or drugs screening. However, the difficulties to culture and image them in 3D hamper their full adoption by laboratories and compagnies. In the meantime, Light Sheet Fluorescence Microscopy technics (LSFM) have proven to be extremely efficient for 3D imaging of biological samples at various spatial and temporal scales with minimal photo-damaging effects.However, LSFM technics are usually restricted in the number of sample and/or condition that can be probed due to complex sample mounting constraints. To address those questions, we develop in collaboration with V.Viasnoffand G. Grenci teams at MBI (NUS, Singapore) a culture and imaging platform combining microfabricated micro-wells,with a single-objective-based LSFM architecture named soSPIM. This combination allows to standardize and parallelize both the culture and the imaging of complex 3D biological models, paving the way toward the use of spheroids and organoids in multi-conditions screening experiments.
In that perspective,we aim to develop new culture vessels that would allow to transform our culture and imaging platform in a multi-condition one. Those new vessels will have to allow the appropriate and timely delivery of media and chemical compounds into the 3D cultured models. Then, a dedicated process will be implemented to allow the automated monitoring of those different conditions using the soSPIM 3D imaging technology.
Missions
The candidate missions will be:
- to develop a custom fluidics system for media and chemical compounds delivery into multi-wells plates,
- to adapt the fabrication process of the JeWell devices to this multi-well plate format. She/he will also participate to the validation of the multi-conditions systems created performing toxicology assays on 3D biological models developed in the team or by collaborators.
Candidate profile
We seek a motivated, enthusiastic and independent candidate, with a strong expertise in fluidics and automation and showing an interest in biology. Complementary skills in fluorescence microscopy, and/or programming would be appreciated. The candidate will work in an English-speaking environment, in close interactions with biologist in Oncology (BRIC).
Contract
Applicants should send a CV, a motivation letter and contact details for at least two referees to: jean-baptiste.sibarita@u-bordeaux.fr and remi.galland@u-bordeaux.fr
#IMadeIt: Marine Cabillic, a woman scientist who knows how to adapt
Read the portrait of Marine Cabillic in English and French
Marine Cabillic, former research engineer in the team of Jean-Baptiste Sibarita, looks back on her five-six years at IINS. She is now software product manager at ONI (Oxford Nanoimaging) where she is responsible for a cloud software solution. At IINS, she has implemented several software solutions, analysis pipelines, as well as biology and imaging protocols for the automation of super-resolution microscopy by localisation
What is your background?
"I have always been interested in the health field. Thus, throughout my career, I have developed tools for the acquisition and analysis of biological or biomedical images. First, I joined ISEN Brest, an engineering school also has a speciality in biomedical technologies. Then, I did my last year of the master’s degree in a work-study program at Medimaps which is a start-up company specialised in medical imaging software. When I graduated, I joined Jean-Baptiste Sibarita’s team [...] and after two years at the Institute, I did a cifre thesis with Sanofi. This was part of a partnership between Sanofi-Aventis in Paris and the Institute. Then, at the end of my thesis and six months as a post-doc, I was hired at ONI."
Why did you join IINS?
"I initially had no expertise in neuroscience. However, I had a background in health and biotechnology, with software design. Moreover, the Sibarita team is a technical team where neuroscience is not the main subject of the research project. [...] For me, IINS is very interdisciplinary and innovative, with very varied projects, tools and fields of application! IINS is internationally recognised, which allows for more collaborations and projects. I joined IINS because I wanted to work in research and innovation. [...] At the Institute, in collaboration with Sanofi, I developed an automated method, combining high-content screening (HCS) and super-resolution, capable of screening and quantitatively characterising therapeutic antibodies for immunotherapy applications. The idea was to characterise the organisation and trafficking of antibody/therapeutic receptor pairs at the membrane of T cells in 96-well plates by quantitative single molecule localisation microscopy. This was done using the HCS-SMLM platform but I also worked on single cells using super-resolution multiplane light sheet imaging (soSPIM)."
As a woman in science, what difficulties have you encountered?
"I faced a lot of technical challenges during my thesis. At the beginning, being quite isolated in terms of a project that was not neuro-focused, I did not dare to go and ask other teams for advice. Also, when the project was not moving in the right direction, I found it difficult to confront the ideas of my collaborators. But the scientific world is very collaborative, to evolve in this world, you have to open up and exchange with others. So this what I did, I openned-up and ask for help of my peers to overcome my difficulties!"
Why do women need to be more recognised in the scientific community?
"In my field, which is technical, there are very few women. So it is more difficult to get ahead. In order to achieve gender equality, it is important to promote more women who carry out research projects or technical theses in order to reach young women as early as possible. Nevertheless, year after year, I notice an increase in the number of women working in technical research!"
Professional pride?
"I have very often embarked on new projects and fields. So I had to learn to adapt. During my thesis with Sanofi, I had to learn everything about biology. And I succeeded, I ended up developing protocols for culture, transfections, immune-labelling and imaging. Before that I had never touched a pipetboy in my life. When I accepted my current position at ONI, I also threw myself into the unknown, a position of responsibility, no initial skills, in English and working from home, big challenge but I adapted!"
Any advice for young researcher?
"Go for it. Why not yourself? First author of a paper, give a talk at that conference, win that prize, try a project, or apply for that kind of job,... Have fun. Science is fun, try projects/collabs, experiments, explore avenues. Train as much as you can and attend conferences,... Get organised. Planning your research projects is a key and important point that will avoid surprises and allow you to change your strategy if necessary and at the right time. Popularise your science! The first few times are hard but you will come out better."
Job offer: engineer to develop light-sheet microscopy for neurosciences applications
Project description
Light Sheet Fluorescence Microscopy technics (LSFM) has demonstrated to be a method of choice for 3D imaging biological samples at various spatial and temporal scales with minimal photo-damaging effects.Several solutions have been developed in the field of neuroscience to image biological samples ranging from fixed whole brains,to single dissociated neurons growing on a coverslip. In this regard, the Interdisciplinary Institute for Neuroscience (IINS)and the Bordeaux Imaging Center (BIC)are equipped with 3complementary LSFMtechniques: (1) an ultramicroscope for whole brain imaging; (2) a Lattice Light Sheet Microscope(LLSM)to image the first layers of brain slices at high spatial resolution; (3) a single objective selective plane illumination microscope (soSPIM)dedicated to 3D cell cultures and in-depth single-molecule localization microscopy (SMLM).
We aim to complete our catalog by implementing a solution based on the Oblique Plane Microscopy (OPM) architecture, which will be dedicated to fast neuronal sample imaging, ie. brain slices,equipped with local photo-manipulation.
Missions
The candidate missions will be:
- to assemble a custom OPM to address specific neurobiological questions,
- to develop the instrument control software and iii) to participate in the testing and optimization of the OPM in close collaboration with neuroscientists.
Candidate profile
We seek for an independent, motivated and enthusiastic candidate, with an interest in neuroscience and a strong expertise in optics and fluorescence microscopy. Strong interest and good skills in programming for instrumentation are required.The candidate will work in an English-speaking environment, in close interactions with the neuroscientists’ team of the Bordeaux’s Neurocampus.
Environment
The candidate will be hosted in the Quantitative Imaging of the Cell team, a R&D team with an internationally-recognized expertise live cell microscopy and quantitative analysis.The Bordeaux Imaging Center, where the final instrument will be eventually transferred,is an imaging platform with a department dedicated to photonic microscopy for biology. It is equipped with several advanced fluorescence microscopy systems (Confocal, STED, SMLM, LLSM, ...). The BIC and IINS are hosted in the same building, a brand new neuroscience research center located on the Carreire campus of the Bordeaux University.
The Interdisciplinary Institute for Neuroscience (IINS) is an international level research center in neurosciences. It gathers 14 teams with complementary and interdisciplinary expertise, as well as several platforms to address cutting-edge questions in various aspects of neurosciences.
Contract
A 1 year, renewable, engineer position is available in the framework of the French “Grands Programmes de Recherche” BRAIN awarded to the Bordeaux Neurocampus. Applicants should send a CV, a motivation letter and contact details for at least two referees to:
jean-baptiste.sibarita@u-bordeaux.fr; remi.galland@u-bordeaux.fr; mathieu.ducros@u-bordeaux.fr
Selected Publications
MORE
MORE
MORE
MORE
MORE
MORE
MORE
MORE
MORE
MORE
MORE
MORE
Members
« Researcher »
| GALLAND Remi | Researcher | remi.galland@u-bordeaux.fr | +33533514748 |  |
| THOUMINE Olivier | Researcher | olivier.thoumine@u-bordeaux.fr | +33533514704 |  |
« Technical Staff »
| BRUGIERE Thibault | Technical staff | thibault.brugiere@u-bordeaux.fr | +33533514700 |  |
| CLOATRE Tiffany | Technical staff | tiffany.cloatre@u-bordeaux.fr | +33533514700 |  |
| LEVET Florian | Technical staff | florian.levet@inserm.fr | +33533514747 |  |
| MONSEIGNE Thibaut | Technical staff | thibaut.monseigne@u-bordeaux.fr | +33533514700 | |
| SIBARITA Jean-Baptiste | Technical staff | jean-baptiste.sibarita@u-bordeaux.fr | +33533514706 |  |
« PhD student »
| BETTAREL Laetitia | PhD student | laetitia.bettarel@u-bordeaux.fr | +33533514700 | |
| IDRISSI Ihssane | PhD student | ihssane.idrissi@u-bordeaux.fr | +33533514700 | |
| NEUHAUS Abdelghani | PhD student | abdelghani.neuhaus@u-bordeaux.fr | +33533514700 |